Genomics of the Chilean Population: genetic characterization necessary for biomedical research, public health, and forensic medicine
Project Summary
Although historical records tell us that the Chilean population was founded by the admixture of local Amerindian populations and colonizing Spaniards, the actual ancestral composition of the genome of Chileans has not been systematically studied. In reality, it is expected the genomic make up of current Chileans is much more complex, with multiple ethnic groups contributing at different times since the initial contact between Americans and Europeans. The general objective is to characterize the ancestral genetic composition of the Chilean population using ancestry informative markers genome-wide in a panel of 500 individuals. In addition, we will select 100 informative SNPs to genotype 2500 individuals from 12 city districts covering the most inhabited regions of Chile. This study will allow a detailed description of the genomic composition of Chileans and it will help in improving our understanding of their genetic heritage. This information will be critical for designing biomedical research protocols that accounts for the diversity of the Chilean population.
Funding
Project funded by FONDEF grant D10I1007. 2012-2014: “Genomics of the Chilean Population: genetic characterization necessary for biomedical research, public health, and forensic medicine”.
Objectives
- Identify a panel of Ancestry Informative Markers that can distinguish between Amerindian and European ancestry genome-wide in Chileans.
- Characterization of the Amerindian/European composition of the Chilean population by geographic location and socioeconomic variables.
Sampling
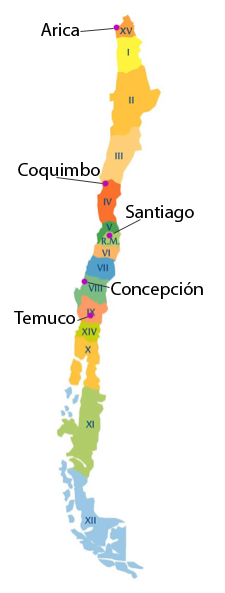
DNA will be obtained from blood donors in each of five cities in Chile: Arica, Coquimbo, Santiago, Concepción and Temuco. In each city, 250 individuals from each of two hospitals will be recruited. In Santiago, we will sample 2 additional hospitals, providing a total of 3000 individuals. Each person will be provided an informed-consent form describing the objective and nature of the study as well as a survey with demographic and socioeconomic questions.
SNP discovery
Among the 3000 individuals, 30 will be selected with 2-4 last names of indigenous origin, 15 Mapuche and 15 Aymara. Their DNA will be sequenced by NGS technology to discover SNPs and short indels not present in dbSNP. This information plus the information available in public databases will be used to construct a panel of 1500 Ancestry Informative Markers (AIMs), i.e. polymorphisms with large differences in gene frequencies between Amerindians and Europeans.
High-density genotyping
We will design a custom genotyping array or select a commercially available platform with representation of the AIMs above selected or corresponding TAG SNPs in close proximity. A panel of at 1500-800K SNPs will be genotyped in a sample of 500 individuals. This information will be used to estimate relevant parameters to describe the level of admixture of Chileans, such as average ancestry percentage, linkage disequilibrium, number of generations since admixture, and number of haplotypes that are private or in common with populations in other regions of the world.
Low-density genotyping in the larger sample
A panel of 100-150 SNPs will be selected from the high-density panel that maximizes ancestry informativeness and genome coverage. This panel will be genotyped by a low-throughput technology, e.g. qPCR, in the remaining 2500 individuals. This will allow characterizing the diversity of ancestry and genomic composition of Chileans across regions, cities, and socioeconomic variables.


